M. Radha¹, M. Nirmala Devi², S. Vishnu Shankar³, P. Jeyalakshmi⁴*
¹ Assistant Professor (Statistics), Department of Agricultural Economics, Anbil Dharmalingam Agricultural College and Research Institute, Trichy, Tamil Nadu, India
*Corresponding author: radha@tnau.ac.in
² Assistant Professor (Statistics), Department of Physical Science and Information Technology, Tamil Nadu Agricultural University, Coimbatore, Tamil Nadu, India
³ Teaching Assistant (Agricultural Statistics), Department of Physical Science and Information Technology, Tamil Nadu Agricultural University, Coimbatore, Tamil Nadu, India
⁴ Assistant Professor (English), Department of Agricultural Extension and Communication, VOCAC&RI, Killikulam, Tamil Nadu, India
Email: jeyalakshmi.p@tnau.ac.in
Introduction
Genetic variability forms the cornerstone of crop improvement and is fundamental to the success of plant breeding programs. Assessing variability among genotypes enables breeders to quantify the extent of differences for economically important traits, thereby identifying heritable components and predicting response to selection. Effective utilization of this variability enhances yield, stress tolerance, quality attributes, and adaptability across diverse environments.
Since observed phenotypic expression is influenced by both genetic makeup and environmental conditions, a major objective of variability studies is to partition total variation into genetic and environmental components. Such partitioning allows the identification of genotypes with stable performance, thereby strengthening breeding strategies.
Components of Variability
Variability in quantitative traits can be separated into the following components:
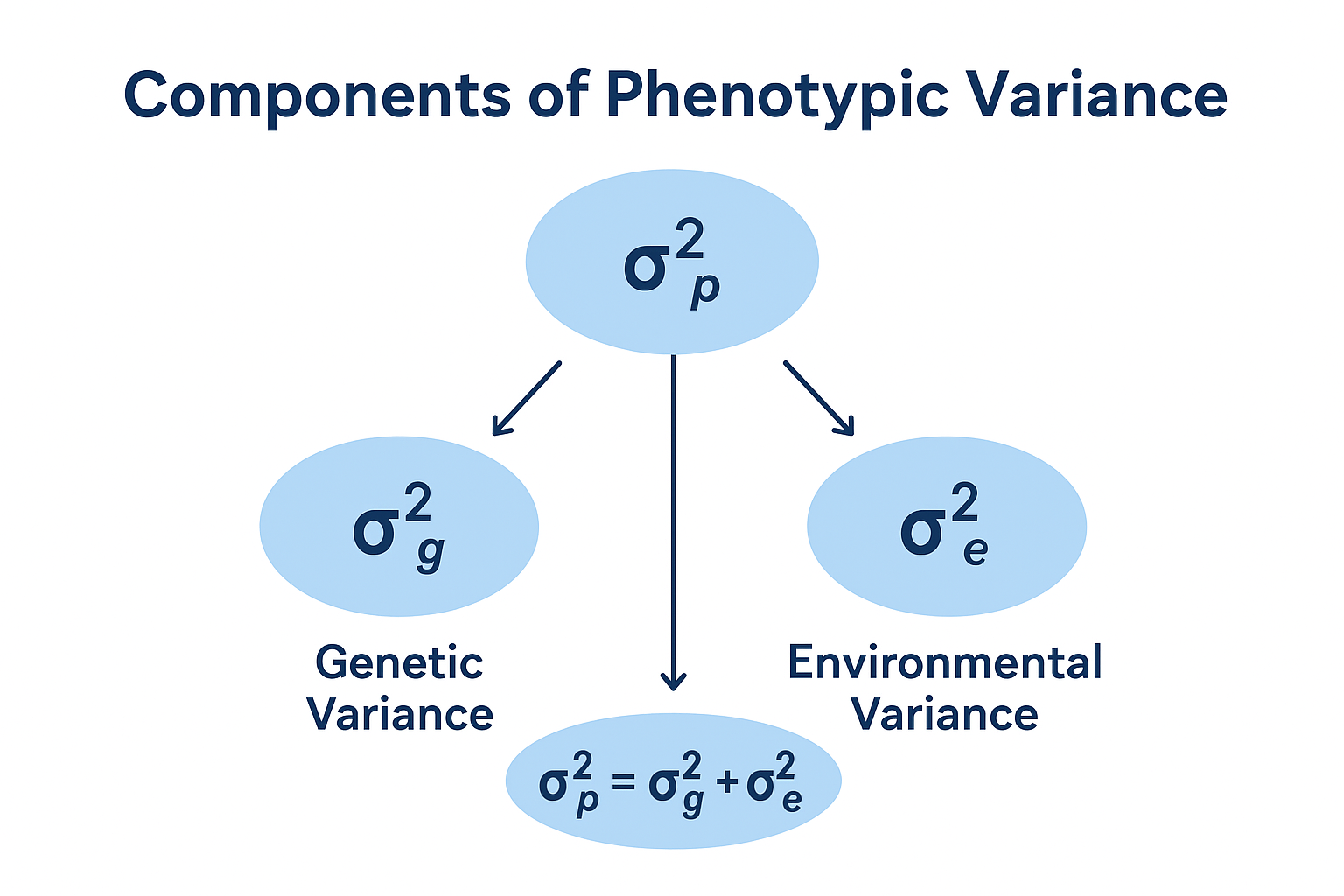
Genotypic Variance (σ²g): Reflects variability due to inherent genetic differences, representing the heritable fraction that breeders seek to exploit.
Environmental Variance (σ²e): Captures the influence of non-genetic factors such as soil fertility, climatic conditions, and management practices.
Phenotypic Variance (σ²p): Denotes the total observable variability, which is the sum of genetic and environmental variances (σ²p = σ²g + σ²e).
These components are generally estimated through analysis of variance (ANOVA), wherein genotypes are considered either fixed or random depending on experimental design.
Genetic Parameters in Variability Analysis
The efficiency of selection in breeding programs is often evaluated using genetic parameters derived from variance estimates:
Genotypic and Phenotypic Coefficients of Variation (GCV and PCV):
These express variability relative to the mean of the trait. A small difference between GCV and PCV suggests minimal environmental influence, whereas a larger difference indicates significant environmental effects.Heritability (h²):
Defined as the proportion of genotypic variance to phenotypic variance. High heritability signifies that phenotypic selection will be reliable, as most observed variation is genetically controlled. Estimates may be broad-sense (H²) or narrow-sense (h²n), depending on whether dominance and epistatic effects are included.Genetic Advance (GA):
Represents the expected improvement in trait performance from one cycle of selection. Expressed as a percentage of the mean (Genetic Advance as Percent of Mean, GAM), high GA coupled with high heritability indicates predominance of additive gene action, thereby favoring direct selection.
Statistical Approaches to Variability Analysis
Analysis of Variance (ANOVA): Widely employed to detect significant differences among genotypes, with mean squares partitioned into genotypic and error components to estimate variance.
Multivariate Approaches:
Principal Component Analysis (PCA): Identifies traits contributing most to variability.
Cluster Analysis: Groups genotypes into distinct classes, facilitating diversity assessment and parental selection.
Molecular Marker-Based Analysis:
Advances in molecular genetics have enabled assessment of variability using markers such as SSRs, SNPs, and AFLPs. Marker-based studies provide precise estimates of genetic relationships, particularly useful where environmental effects confound phenotypic assessment.
Relevance of Variability Analysis in Breeding Programs
Identification of Superior Genotypes: Facilitates screening for desirable trait combinations.
Breeding for Stability: Multi-environment variability estimates help in selecting genotypes with wide adaptability.
Genetic Conservation: Knowledge of variability guides conservation of diverse and unique germplasm.
Formulation of Crop Improvement Strategies: Determines whether direct selection, hybridization, or advanced breeding techniques are most appropriate.
For instance, in self-pollinated crops such as rice and wheat, high heritability coupled with high genetic advance favors direct selection. Conversely, in cross-pollinated crops like maize, where dominance and epistatic effects are more pronounced, hybrid breeding is often more effective.
Conclusion
Quantitative analysis of variability provides critical insights into the genetic architecture of crop traits and their potential for improvement. By distinguishing between genetic and environmental sources of variation, breeders can implement targeted selection strategies to achieve sustainable gains. The integration of classical biometrical tools with molecular markers and advanced statistical approaches enhances the accuracy of variability estimates, thereby accelerating crop improvement to meet the challenges of climate change and global food security.
References
Burton, G. W., & De Vane, E. H. (1953). Estimating heritability in tall fescue (Festuca arundinacea) from replicated clonal material. Agronomy Journal, 45(10), 478–481.
Falconer, D. S., & Mackay, T. F. C. (1996). Introduction to Quantitative Genetics (4th ed.). Longman.
Johnson, H. W., Robinson, H. F., & Comstock, R. E. (1955). Estimates of genetic and environmental variability in soybeans. Agronomy Journal, 47(7), 314–318.
Allard, R. W. (1999). Principles of Plant Breeding (2nd ed.). Wiley.
Hallauer, A. R., Carena, M. J., & Miranda Filho, J. B. (2010). Quantitative Genetics in Maize Breeding. Springer.
Singh, P., & Chaudhary, B. D. (1985). Biometrical Methods in Quantitative Genetic Analysis. Kalyani Publishers.